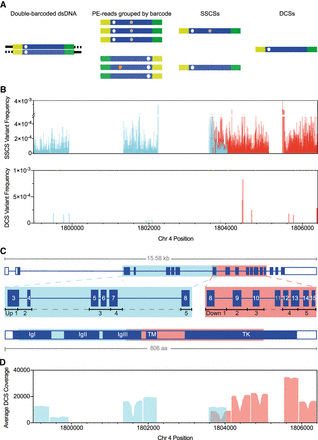
DS using FGFR3 targeted sequencing strategy with enzymatic digestion. (A) DS overview: Barcoded adapters (yellow and green) were ligated at both ends of size-selected restriction enzyme–digested genomic fragments (blue). After several rounds of PCR amplification and hybridization captures, libraries were sequenced on the Illumina MiSeq (v3 600-cycles); for sequence analysis, paired-end (PE) reads were grouped into SSCSs, and complementary SSCS were joined into a DCS. Only DCS with mutations in both complementary SSCS (white asterisk) were considered true variants. Colored asterisks represent artifacts (e.g., sequencing and PCR errors). (B) Variants detected at 2881 different positions of the total 4405 sequenced positions identified in SSCSs (n = 14,552 total variants; VAF = 1.0 × 10−5 to 3.8 × 10−3) or at 15 positions in DCSs (n = 15 total variants; VAF = 6.0 × 10−5 to 8.3 × 10−4) of two FGFR3 libraries (FGFR3 Up O Oct19 Re-seq and FGFR3 Down O BAT), each targeting either the Up 1-5 (blue) or the Down 1-5 (red) regions/subregions. (C) Exonic structure of FGFR3. Shown are the five upstream (blue) or downstream (red) regions targeted via restriction enzyme fragmentation that include exons 3 to 8 or 8 to 15 (except exon 11), respectively; restriction digests rendered fragments ∼370–550 bp in size. Acronyms of the FGFR3 structure are as follows: (IgI-III) immunoglobulin-like domain I-III, (TM) transmembrane domain, (TK) tyrosine kinase domain, and (aa) amino acids. (D) Average DCS coverage per position of all sequenced libraries.











