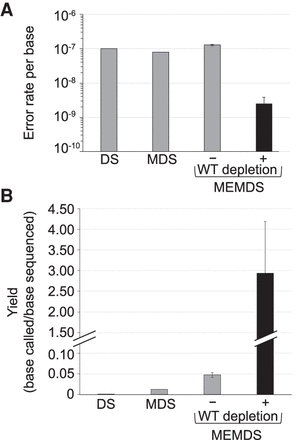
Accuracy and yield of MEMDS compared with current cutting-edge methods for studying target regions. (A) Under a highly conservative estimate, MEMDS increases accuracy by at least 40-fold compared to duplex sequencing (DS) (Kennedy et al. 2014) and maximum depth sequencing (MDS) (Jee et al. 2016). (B) MEMDS also increases yield per sequenced base (i.e., the number of MEMDS confirmed bases divided by the number of paired-end sequenced bases) by orders of magnitude over both DS and MDS (Kennedy et al. 2014; Jee et al. 2016). Notice that in MEMDS, the yield can be higher than 1 because the mutation enrichment factor is accurately calculated (Supplemental Text S2) and the base identity is known for the ROI sequences that were digested and removed from the final sequencing libraries (they have the restriction enzyme recognition sequence). Although the accuracy of DS has been improved in the context of sequencing large parts of the genome (Abascal et al. 2021), yield considerations and other limitations preclude applying current DS-based methods to narrow ROIs and target mutations (Kennedy et al. 2014; Supplemental Text S1) with the same efficiency as that of MEMDS.











