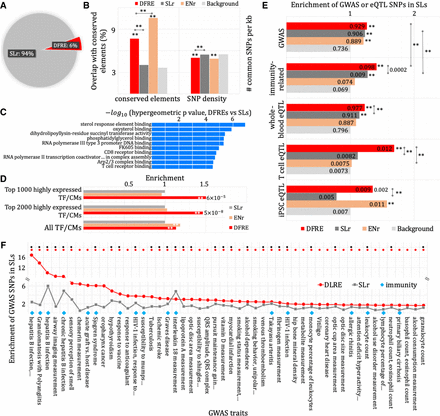
DFREs are consequential. (A) A distribution pie chart of the silencers (SLrs) and DFREs for primary T cells in peripheral blood. (B) Evolutionary sequence conservation of DFREs (red), SLrs (gray), and ENrs (orange) computed using the overlap with conserved segments and the density of common SNPs. The asterisks above the bars indicate the significant enrichments compared with the background consisting of a group of the randomly sampled genomic regions with the GC and repeat contents matching the silencers. (**) P < 10−10. (C) Molecular functions significantly associated with the DFREs. The results are from the web tool GREAT using all silencers as background. (D) Enrichment of DFREs in the loci of the TFs and CM genes. (**) P < 10−5. White asterisks in the bars represent significant enrichment compared with the SLrs. The presented P-values are the enrichments compared with all TF and CMs. (E) Enrichment of GWAS SNPs and eQTLs in DFREs, SLrs, and ENrs. The density of SNPs per kilobase is listed in the bars. The asterisks (and the values) next to the bars quantify the significance of enrichment compared with the background. (**) P < 10−5. (F) GWAS traits with which the associated SNPs are enriched in the DFREs compared with ENrs. Only the GWAS traits significantly enriched in the DFREs are presented. The azure diamonds highlight immunity-related traits. Red and black dots on the top indicate the significant enrichments (P < 0.05) of GWAS SNPs in DFREs and SLrs, respectively. The details about these results are listed in Supplemental Table S2.











