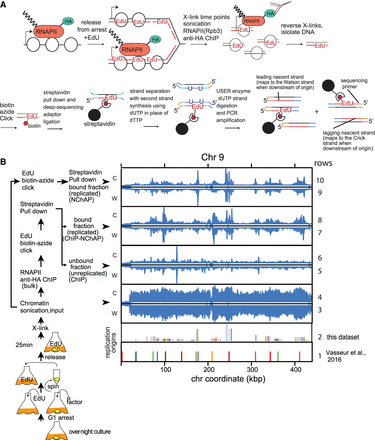
RNAPII ChIP-NChAP in early S phase. (A) Diagram of the RNAPII ChIP-NChAP experiment. (B) RNAPII distribution on Chromosome 9 from chromatin fractions diagrammed on the left, in early S phase 25 min after release from G1 arrest. The positions of replication origins are shown in rows 1 and 2—(1) identified in Vasseur et al. (2016); and (2) from this data set. Read counts from all fractions were grouped in 50-bp bins and first normalized to the genome average read count and then to the highest peak value in each chromosome. W and C are reads from Watson and Crick strands, respectively.











