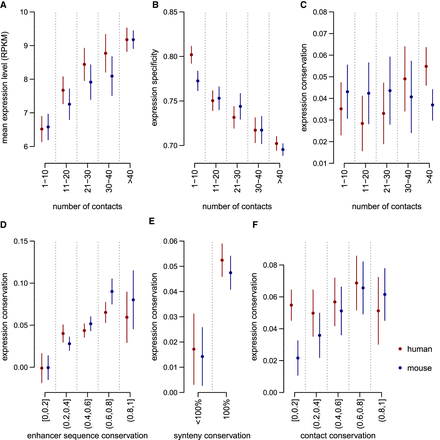
The complexity and the evolution of cis-regulatory landscapes are associated with gene expression evolution. (A) Average expression levels as a function of the number of contacted enhancers in PCHi-C data for human (red) and mouse (blue). (B) Gene expression specificity index (tau, which ranges from zero for housekeeping genes to one for highly specific genes; Methods) as a function of the number of contacted enhancers. (C) Expression conservation as a function of the number of contacted enhancers in PCHi-C data. (D) Expression conservation as a function of the average sequence conservation of contacted ENCODE enhancers. (E) Expression conservation, depending on whether or not genes underwent at least one break of synteny with the contacted enhancers between human and mouse genomes. (F) Expression conservation as a function of the proportion of promoter–enhancer contacts conserved in the other species’ PCHi-C data. (A–F) Dots represent median values across all genes in a class; vertical segments represent 95% confidence intervals for the median. (C–F) Gene expression conservation is measured with a Spearman's correlation coefficient between human and mouse relative expression profiles, for pairs of one-to-one orthologous genes, across organs and developmental stages (expression data from Cardoso-Moreira et al. 2019). Expression conservation is further corrected to account for the effect of expression levels and of expression specificity with a multiple linear regression model (Methods). Enhancer predictions are taken from ENCODE. Expression conservation values are the same for both species, but PCHi-C contact maps differ.











