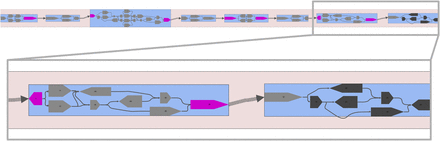
Multiplex de Bruijn graph produced by LJA for CAMP's smoothed and virtual reads. We generated smoothed reads based on mutations identified by NaiveFreq at p = 1%. This is a MetagenomeScope visualization (Fedarko et al. 2017) of the GFA file produced by LJA. Gray pentagons (nodes in this visualization) correspond to “segments” described in this GFA file, which in turn correspond to edges in the de Bruijn graph. Blue regions of the graph indicate “bubble” patterns MetagenomeScope identified in the graph (Miller et al. 2010), highlighted for clarity. Segments colored in pink represent segments that are shared between multiple bubbles: MetagenomeScope duplicates segments (creating a link between the two copies of a duplicate segment) in order to simplify the visualization of adjacent bubbles in the graph. This visualization makes clear that the entire de Bruijn graph can be represented as a linear sequence of bubbles. This topology is consistent with the expected structure of an assembly graph of multiple strains of a genome, for example, as shown in Figure 4 of the work of Kolmogorov et al. (2020); the branching paths in these bubbles likely represent strain-level diversity. The rightmost two bubbles in the graph are shown up in a box below the main drawing of the graph. The five dark-colored segments highlighted in the rightmost bubble correspond to the segments that overlap gene 1217, the most mutated gene in CAMP (Fig. 5, top). There exist three paths through these segments’ bubbles: the top two paths correspond to the “reference” haplotype of gene 1217 and the bottom two paths correspond to the “alternate” haplotype of gene 1217 (Supplemental Material, “Haplotypes of the most mutated gene in CAMP”; Supplemental Fig. S20). The reason for the reference haplotype being represented by two distinct paths is that these paths cover other mutated positions located earlier in CAMP.











