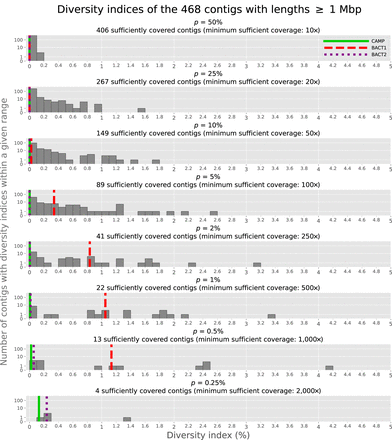
Diversity indices vary widely across the 468 long contigs in SheepGut. Smaller values of p allow us to “call” increasing amounts of p-mutations in a MAG, at the cost of requiring higher sequencing coverage to reliably distinguish these mutations from errors
(we note that this differs from ordinary usage of NaiveFreq, which does not explicitly take coverage into account). The Supplemental Material, “Diversity index details,” provides details about some of the high-diversity-index contigs shown in this figure and about
how minimum sufficient coverages are determined for each value of p (we use minReadNumber = 5 here). The diversity index values for the three selected MAGs are highlighted on each row of the histogram as vertical
lines. Although it is shown in Figure 3, we have omitted p = 0.15% from this figure because its minimum sufficient coverage is 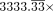 ; the only MAG that is sufficiently covered for this value of p is CAMP.
; the only MAG that is sufficiently covered for this value of p is CAMP.











