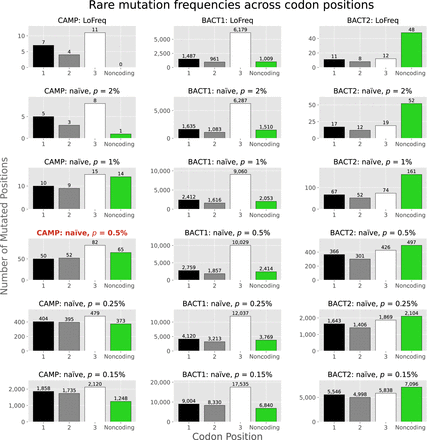
Rare mutation frequencies vary across codon positions in the three selected MAGs. This plot includes variant calls from LoFreq (top row) and results from NaiveFreq for p ∈ {2%, 1%, 0.5%, 0.25%, 0.15%} (bottom rows). The expected “pattern” of codon position mutation frequencies (MCP2 < MCP1 < MCP3) holds for all plots except for the case of CAMP at p = 0.5% (the title for this plot is highlighted in red). As discussed in the Supplemental Material, “Applying LoFreq to the SheepGut dataset” (Supplemental Fig. S7), LoFreq's results are similar to NaiveFreq's at p = 2%. To focus this visualization on “rare” mutations, we limit the mutations included in all rows (including LoFreq's) to those located in positions at which freq(pos) < 5% (Methods). We also treat positions where LoFreq called multiple mutations as if only a single mutation were called at this position (Supplemental Material, “Applying LoFreq to the SheepGut dataset”). The Supplemental Material, “Codon position analysis details” (Supplemental Fig. S9), shows an alternate version of this figure, in which y-axis values are normalized by the total number of positions considered in order to make plots from different MAGs more easily comparable.











