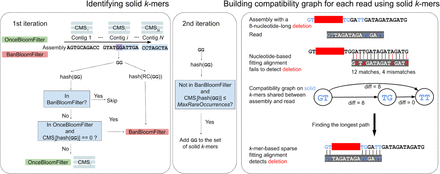
VerityMap pipeline. (Left) The input of VerityMap is an assembly (a set of contigs) and a set of reads that contributed to this assembly. VerityMap iterates through each contig twice in order to identify solid k-mers. At the first iteration, VerityMap stores k-mers that appear in multiple contigs and all reverse-complementary k-mers within the BanBloomFilter. For each contig, VerityMap constructs a CMS that counts occurrences of k-mers within this contig. Finally, VerityMap uses OnceBloomFilter (and BanBloomFilter) to distinguish between rare k-mers that appear within a single and multiple contigs. Both Bloom filters and the CMS corresponding to the current contig are being modified simultaneously during the first iteration through the assembly. At the second iteration, VerityMap queries the constructed data structures to identify the set of solid k-mers. (Right) Aligning a read GTTAGATAGATGGATT against a misassembled contig GTTGGATTGATAGATAGATG with an 8-nucleotide-long deletion TAGATAGA (solid k-mers are shown in blue). The solid k-mer GT (TG) precedes (follows) the deletion breakpoint. The nucleotide-based fitting alignment fails to identify this deletion owing to limitations of the standard scoring approaches in highly repetitive regions. In contrast, VerityMap identifies this deletion using the k-mer-based sparse fitting alignment and a new scoring approach. To achieve this goal, it constructs a compatibility graph on all pairs of solid k-mers shared between a read and the assembly and finds a longest path in this graph. The new scoring reflects the discrepancies in distances between solid k-mers in the assembly (distance 2 between GT and TG in the assembly) and solid k-mers in the read (distance 10 between GT and TG in the read), resulting in diff(GT,TG) = 8. VerityMap incorporates these discrepancies into the edge-weights of the compatibility graph and outputs a longest path in this graph as the primary read alignment.











