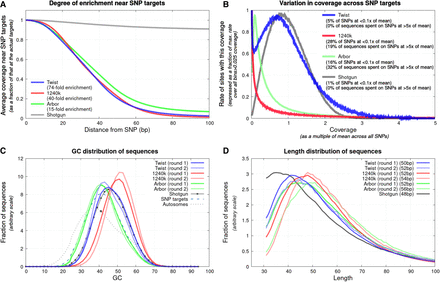
Characterization of enrichment. (A) Degree of enrichment as a function of distance from 1,150,639 targeted autosomal SNPs (position 0) for the 15 high-coverage libraries at the bottom of Table 1; enrichment at the SNP relative to positions 100 bp away is shown in the legend. (B) Variation in coverage across SNP targets for the same libraries. (C) Proportion of nucleotides that are guanine or cytosine (GC) has a downward bias relative to the unenriched library for Arbor, upward for 1240k, and little bias for Twist Ancient DNA; this analysis uses data from the first 10 libraries in Table 1 with full results from both rounds of capture. (D) All assays preferentially enrich longer molecules, with the least length effect for Twist Ancient DNA (medians in legend, 10 libraries of data). All plots reflect data before removal of duplicated sequences as our goal is to study effectiveness of enrichment on a per-molecule basis.











