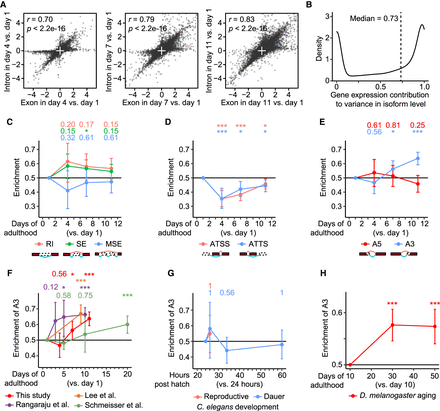
Aging increases the alternative selection of distal 3′ splice sites (A3) in transcript isoforms. (A) Comparisons between age-dependent expression changes of exon and intron regions in individual nonoverlapping genes. Pearson correlation coefficient r and its significance P are marked. (B) Gene expression contribution to variance in isoform level. See Supplemental Figure S2A for the schematic overview. (C–E) Enrichment of different RNA-processing events in isoform fractions that were up-regulated during Caenorhabditis elegans aging. RNA-processing events include retained introns (RI), skipped exons (SE), multiple skipped exons (MSE) (C), alternative transcription start sites (ATSS), alternative transcription termination sites (ATTS) (D), alternative 5′ splice sites (A5), and alternative 3′ splice sites (A3) (E). (F–H) Enrichment of A3 in isoform fractions up-regulated during aging (F), during reproductive and dauer larval development (G) in C. elegans, and during aging in Drosophila melanogaster (H). We noticed that only a single A3 isoform was detected as up-regulated in the comparison between animals at 34 h post-hatching and at 24 h post-hatching in reproductive larval development. Adjusted P value is shown at the top of each panel, calculated by one-proportion Z-test adjusted using false discovery rates; (*) adjusted P < 0.1, (***) adjusted P < 0.001.











