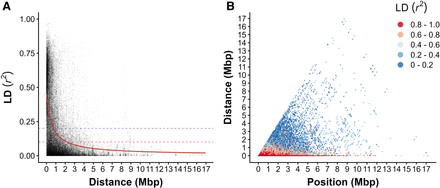
Within-scaffold linkage disequilibrium (LD) decays slowly in WRC. (A) LD was assessed using SNPs with a minor allele frequency (MAF) cutoff of 0.05 to reduce error associated with rare alleles (n = 16,202 SNPs). Decay was estimated using a nonlinear model (red line); r2 decayed to baseline thresholds of 0.2 (purple dotted line) and 0.1 (pink dotted line) at 0.751 and 2.17 Mbp, respectively. (B) Pairwise LD for all pairs of SNPs (n = 16,202). Each point on the plot represents the LD between two SNPs at a given distance from one another and relative position on the scaffold. Color indicates LD range, with red indicating SNPs in strong LD.











