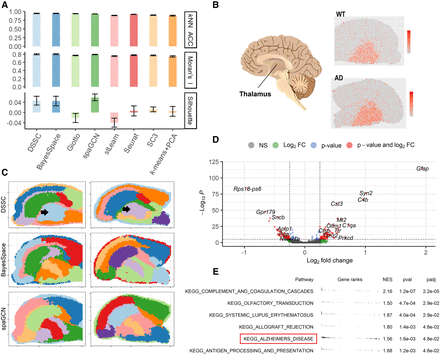
Results of 10xMBAD data sets. (A) Clustering performance (without true labels). (B) Cartoon of the brain showing the position of the thalamus and the expression of a marker gene, Tcf7l2, for the thalamus in a WT and an AD sample. (C) Predicted labels for a WT sample and an Alzheimer's disease (AD) sample from DSSC, BayesSpace, and SpaGCN; the black arrows indicate the thalamus regions. (D) Volcano plot from the differential expression analysis (DE) between the cells in the thalamus from the WT and the AD samples. (E) KEGG pathway analysis from the DE results in D. The pathway of AD is highlighted by the red box.











