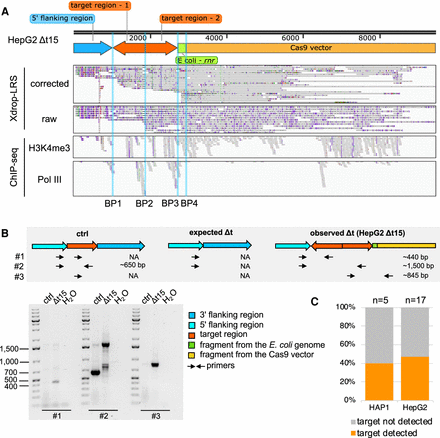
A duplication, inversion of target-derived fragments, and integration of exogenous DNA fragments arose in the deletion clone HepG2 Δt15. (A) The genome browser view shows Xdrop-LRS (top, corrected and raw reads) and ChIP-seq (bottom, histone H3 lysine 4 trimethylation [H3K4me3] and RNA polymerase III [Pol III] reads) aligned against the assembled contig supporting four different breakpoints (BP1-4, blue vertical lines) within the contig. Arrows (top) show genomic orientation and approximate size of the flanking (blue) and target (orange) regions, as well as sequences of the E. coli genome (green) and the CRISPR-Cas9-gRNA-Δt-1 transfection vector (yellow) present within the contig. (B) Schematic illustration of the strategy to validate the contig and the expected PCR product length (top) of alleles with an unmodified control (ctrl), expected and observed editing event in deletion clone HepG2 Δt15. Alleles with the expected deletion and on-target alterations exist in deletion clone HepG2 Δt15. Agarose gel image (bottom) visualizes the size of the PCR products. Marker bands specify DNA size in bp. (C) Stacked bar plot indicates the frequency of on-target genomic alterations in the validated HAP1 (n = 5) and HepG2 (n = 17) Δt deletion clones.











