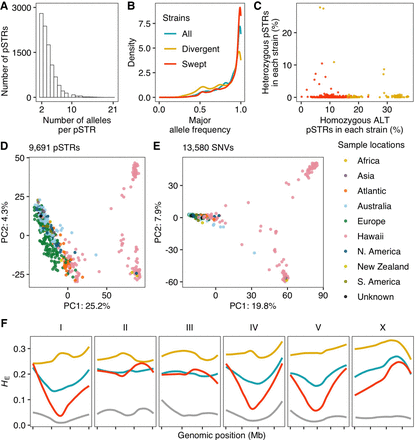
Genetic diversity of Caenorhabditis elegans short tandem repeats (STRs). (A) The distribution of allele counts per polymorphic STR. (B) Major allele frequencies of all polymorphic STRs (pSTRs) for all strains (in blue), divergent strains (in yellow), and swept strains (in red) are shown. (C) The percentage of pSTRs with heterozygous alleles is plotted against the percentage of pSTRs with homozygous alternative (ALT) alleles for each of the 540 strains. Divergent and swept strains are colored yellow and red, respectively. (D,E) Plots show the top two axes of variation, as determined by principal components analysis (PCA) of the genotype covariances using polymorphic STRs (D) and single nucleotide variants (SNVs) (E). Each dot represents a strain and is colored by the sampling location. (F) Chromosomal expected heterozygosity (HE) of pSTRs is shown as locally regressed lines for all strains (in blue), divergent strains (in yellow), and swept strains (in red). Chromosomal HE of SNVs among swept strains is shown as locally regressed lines (in gray). Tick marks on the x-axis denote every 5 Mb.











