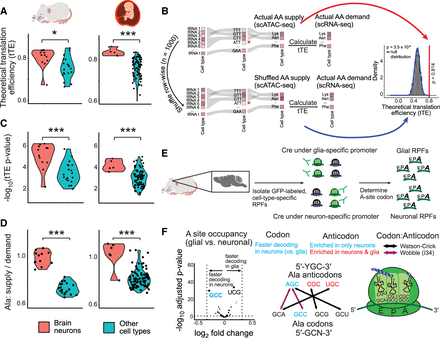
Correlation between AA supply and demand is high and statistically significant in all cell types, with the strongest correlation in brain neurons. (A,C,D) Violin plots are divided into mouse (left) and human (right) showing tTEs (A), tTE P-values (C), and alanine (Ala) supply to demand ratio (D) of brain neurons and other cell types. Of all 20 AAs, only Ala is significantly enriched between brain neurons and all other cell types. (B) Schematic representation of the approach used to determine statistical significance of correlation between AA supply (from tRNA side) and AA demand (from mRNA side), defined here as theoretical translation efficiency (tTE). The observed tRNA expression for each cell type is shuffled 1000 times, pooled at the AA supply level, and correlated to the AA demand (unshuffled) to detect a null distribution of tTE values and determine statistical significance of actual tTE. (E) Workflow from an adult mouse brain ribosome profiling data set (Scheckel et al. 2020). Cell type–specific ribosome-protected fragments (RPFs) for neurons and glia were obtained with cell type–specific GFP-labeling of a ribosomal protein, followed by immunoprecipitation against GFP. The ribosome A-site of these cell type–specific RPFs was determined, and a differential analysis was performed using DESeq2. (F) Volcano plot shows enrichment of tRNAAla (AGC) anticodon in neurons, and faster decoding of the tRNAAla (GCC) anticodon is observed in neurons compared with glial cells. AGC must decode GCC via an adenosine to inosine modification at the first anticodon position. Asterisks display degree of significance: (*) P < 0.05, (**) P < 0.01, (***) P < 0.001; Mann–Whitney U test.











