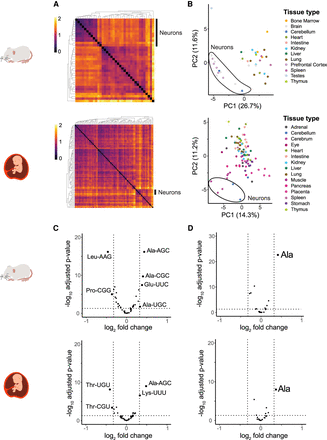
Anticodon usage is also similar across cell types, with brain neurons clustering separately. (A) Heatmaps show the Euclidean distance between anticodon usage across cell types. Only cell types with more than 5000 scATAC-seq cuts were analyzed (see Fig. 3D). The neuronal cluster is indicated with a black line on the right side of the heatmap. (B) PCA plots separate anticodon usage across cell types. The brain neuron cluster is indicated. (C,D) Volcano plots display differences in anticodon usage (C) and AA supply (D) between brain neurons and all other cell types (−log10 adjusted P-values and log2 fold change [FC] as determined using DESeq2). Vertical lines indicate a FC >25%. All panels: mouse (top), human (bottom).











