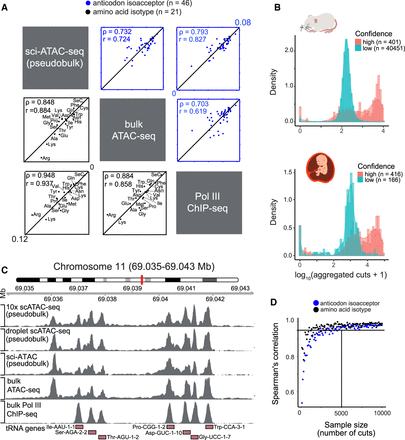
Single-cell ATAC-sequencing is robust for measuring tRNA gene usage. (A) Scatter plots correlate mouse brain scATAC-seq (aggregated in pseudobulk), bulk ATAC-seq, and Pol III ChIP-seq data sets on the anticodon isoacceptor (blue) and AA isotype (black) level. Spearman's rank (ρ) and Pearson (r) correlation coefficients are indicated on the top left corners. (B) Density plots (in log-scale) show the total number of cuts from aggregated scATAC-seq data from the mouse and human scATAC-seq atlases, based on confidence predicted from tRNAscan-SE. (C) Genome browser view illustrates a mouse tRNA gene cluster on Chromosome 11. The location of tRNA genes is shown at the bottom, including upstream and downstream 100-bp flanking regions. Different tRNA genes, including those as close as 220 bp apart (Asp-GUC-1-10 and Gly-UCC-1-7), have distinct peaks in three scATAC-seq data sets (pseudobulk), a bulk ATAC-seq data set, and a Pol ChIP-seq data set (all mouse brain). The leftmost peak present in all ATAC-seq data sets but absent in the Pol III ChIP-seq data set corresponds to the promoter of protein-coding gene Ctc1. This peak is ignored when quantifying tRNA gene usage because it falls outside of 100-bp flanking region of a tRNA gene. (D) Scatter plot determines how many total cuts are needed after pooling cells of the same cell type to obtain reliable information of anticodon isoacceptor and AA isotype usage. A sample size of 5000 cuts per cell type consistently yielded a Spearman's rank correlation coefficient >0.95 to the aggregated scATAC-seq data from mouse and human.











