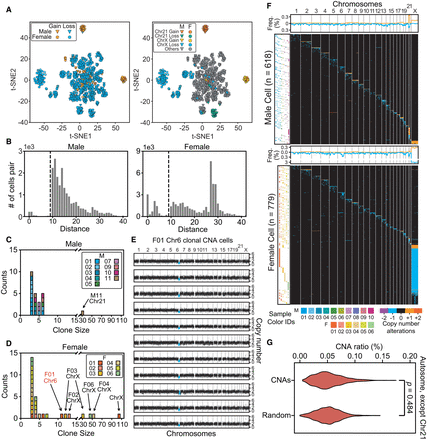
The landscape of cells with copy number alterations. (A) Low-dimensional representations produced through multidimensional scaling of the copy number profiles for cells with >10-Mb CNAs. Colors label different types and locations of CNAs. t-SNE, t-distributed stochastic neighbor embedding. (B) The distributions of Euclidean distances between cell pairs. Two clusters were clearly separated by distance (d) <10 (indicated by dashed lines) in both male and female samples. (C,D) The sizes (number of cells in a clone) and counts of clonal CNAs. Each block in the bar plot represents a clone. Most clonal CNAs with bigger clone sizes were on Chr 21 and Chr X, although there are some small clones with cell numbers of ∼3–5. (E) Copy number profiles of each clonal CNA on Chr 6 in 11 cells from F01. Each graph represents one cell and the blue dots represent the regions with copy number alterations. (F) Overview showing every cell with >10-Mb CNAs each. The heat map (in black) demonstrates the genome patterns for cells with CNAs, which are labeled by different colors for gain and loss. Cells were sorted according to the chromosome fraction carrying CNAs. Each row represents one cell. The scatterplot (left) shows the cells’ sample IDs. The density curve along the top of each heat map shows the aggregate frequency, at 200-kb bins, among all samples for each genomic locus. (G) CNA frequency distribution for each genomic locus in all autosomes except for Chr 21. The distributions of this study's CNAs and of randomly generated CNAs were not significantly different (Mann–Whitney U test), thus indicating a random generation mechanism.











