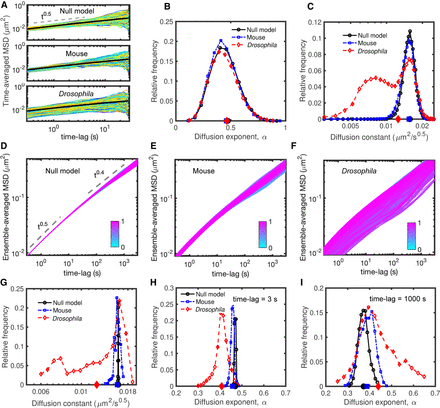
Dynamic properties of simulated chromosomes. (A) Time-averaged MSDi,c of single trajectories of length 30 sec sampled every 0.3 sec for null (upper), mouse (middle), and Drosophila (bottom) models. (B) Distribution of the diffusion exponents αi,c extracted from the time-averaged MSD curves given in A. (C) Distribution of the diffusion constants Di,c for the time-averaged MSDs in A having an exponent αi,c ≈ 0.5. Distributions for other αi,c values are given in Supplemental Figure S3. In B and C, average values of the distributions are shown on the horizontal x-axis. (D–F) Ensemble-averaged (over all trajectories) MSDi of all genomic loci for null (D), mouse (E), and Drosophila (F) models. To discard trivial positional effects, we exclude the last 50 monomers at the two ends of the polymers. Individual ensemble-averaged MSDs were colored from cyan (first monomer) to magenta (last monomer). (G–I) The distributions of the diffusion constant Di (G) and of the diffusion exponent αi (Δt) for short (Δt = 3 sec) (H), and large (Δt = 1000 sec) (I) time scales, extracted from ensemble-averaged MSD curves in panels D through F. The average values of the distributions are shown on the horizontal axes.











