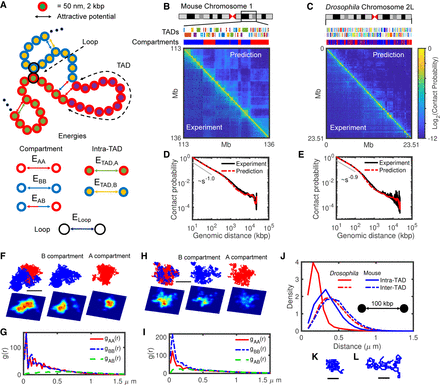
The heteropolymer model and comparison with Hi-C maps. (A) Schematic representation of the heteropolymer model with different structural components and their associated interactions. Each monomer hosts 2 kbp of DNA, and its diameter is 50 nm. The surrounding red (blue) color indicates A (B) compartment, inner different colors correspond to different TADs, and loop anchors are shown by black outer circles. All six possible attractive interactions are shown by arrows in this cartoon. (B,C) Visual comparison of predicted Hi-C maps and the experimental ones for 23 Mbp of mouse Chromosome 1 (113–136 Mbp) and Drosophila Chromosome 2L. The upper tracks show the corresponding TADs and compartments for each chromosome. (D,E) Contact probabilities extracted from predicted (red) and experimental (black) Hi-C maps shown in B and C, respectively. (F,H) Typical snapshots for mouse Chr 1 (F) and Drosophila Chr 2L (H) in mixed and separate A (red) and B (blue) compartments, with corresponding 2D density plots (the lower panels) with axial resolution of 1200 nm, lateral resolution of 100 nm, and pixel size of 50 nm. Bars represent 1 μm of real space. (G,I) Radial distribution functions (g) between A-A (red), B-B (blue), and A-B (green) monomers for mouse (G) and Drosophila (I). (J) Comparison between intra- (full curves) and inter-TAD (dashed curves) distance distributions of pair monomers separated by 100 kbp along the genome for Drosophila (red) and mouse (blue). (K,L) Typical snapshots of ∼300-kbp-long TADs for Drosophila (K) and mouse (L). Bars represent 0.25 μm of real space.











