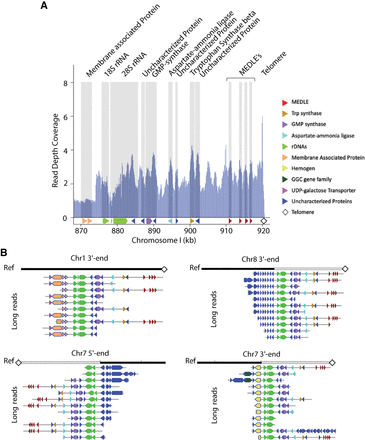
Resolution of repetitive subtelomeric regions found on Chr 1 identifies missing telomeres on Chromosomes 7 and 8. (A) Illumina reads from CpIA are mapped to the CpIA Chr 1 long-read assembly subtelomeric region to identify read pileups and estimates of sequence copy number by normalizing against the average genomics Illumina read depth. Vertical gray areas indicate regions with annotated genes. Annotated genes are represented below the shaded regions, the 5.8S rRNA is present but not indicated. (B) Subtelomeric variaton observed on different CpIA chromosomes is supported by CpBGF ONT long reads. Individual ONT long reads provide evidence of at least four different yet related subtelomeric regions that extend into the chromosomes that were missing telomeres in CpIA (Chr 7 and Chr 8) in addition to Chr 1. The white and black reference bar above each collection of annotated ONT reads identify the resolved subtelomeric regions (white) and linkage to existing assembly (black). The penultimate read on the Chr 7 3′ end panel indicates a unique region of insertion (nucleotide positions 1,191,705–1,217,462). This region contains mostly uncharacterized proteins and two transferases. Each ONT read is annotated as indicated in the key shown in A.











