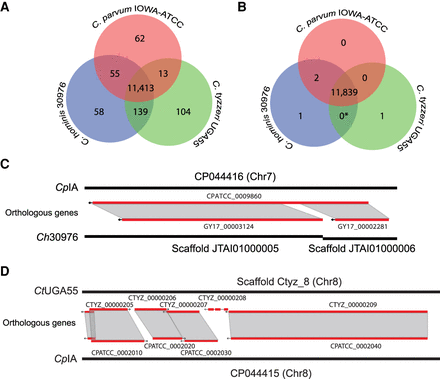
Ortholog distribution of protein-coding genes reveals few differences between the species. (A) Venn diagram of automated protein orthology assignment between CpIA, Ch30976, and CtUGA55. (B) Venn diagram of the same orthologous genes following manual investigation and removal of putative false positives. (*) The 139 genes shared between C. hominis and C. tyzzeri in A are in complex regions with repeats and gaps and do not have enough evidence to prove their uniqueness at this stage (Supplemental Table S7). (C) Example of a false positive paralog count caused by gene fragments on different scaffolds. (D) Putative unique uncharacterized gene found on Chr 8 in CtUGA55.











