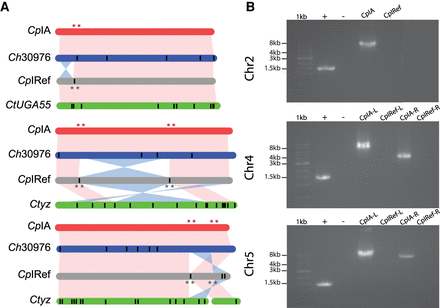
Syntenic relationships between select Cryptosporidium chromosome assemblies. (A) Synteny between Chromosomes 2, 4, and 5. Vertical black lines within a chromosome represent known physical gaps. Synteny between chromosomes is shown in pink and inversions in blue. (B) PCR validation using C. parvum KSU-1 DNA (Supplemental Table S4). Lanes 1, 2, and 3 in all gels are 1-kb ladder, positive control Dnmt2 gene, and no template control, respectively. The remaining lanes test each orientation of the left (L) and right (R) inversion boundaries. Red stars indicate the location of primers designed based on the CpIA assembly, and gray stars indicate the same on the CpIRef assembly.











