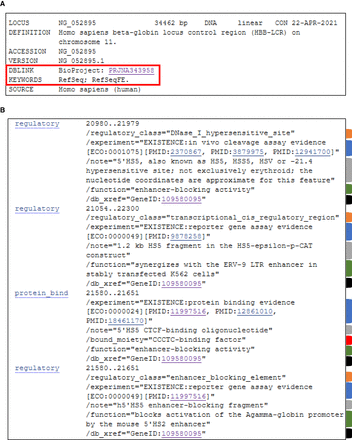
Example of a biological region RefSeqFE flat file. Segments of RefSeq accession NG_052895.1 representing the hemoglobin subunit beta locus control region (HBB-LCR) are shown. (A) Top section of the flat file with a link to BioProject accession PRJNA343958 and the “RefSeqFE” keyword outlined in red. (B) Segment of the feature annotation section. Features are displayed for the 5′HS5 DNase I hypersensitive site (Tuan et al. 1985; Dhar et al. 1990; Wai et al. 2003), a transcriptional cis-regulatory region (Long et al. 1998), a CTCF binding site (Farrell et al. 2002; Bulger et al. 2003; Chan et al. 2008), and an enhancer-blocking element (Farrell et al. 2002). Features include “/experiment” qualifiers with experimental evidence from the literature as indicated by ECO strings and IDs and links to publications (blue tabs), “/note” qualifiers with descriptive information (gray tabs), “/function” qualifiers describing the function of each feature where applicable (green tabs), and a “/bound_moiety” qualifier for the protein-binding site (red tab). All features include a “/db_xref” qualifier (black tabs) linking to the biological region record in the Gene database (GeneID:109580095), and an INSDC class qualifier when relevant (orange tabs).











