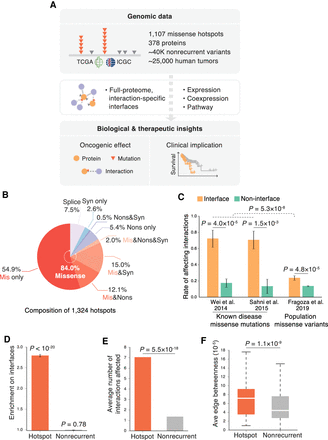
Proteome-wide structural analysis of mutational hotspots. (A) Data resource of mutational hotspots and workflow of our full-proteome interaction-specific characterization framework. (B) Composition of mutational hotspots collected in this study. (Mis) Missense, (Nons) nonsense, (Syn) synonymous, (Splice) splice site. (C) Fraction of missense mutations affecting protein interactions from published experiments. The error bars indicate standard error of the fraction. (D) Distribution of hotspots and nonrecurrent variants on proteins with regard to protein interaction interfaces. Enrichment was calculated as the ratio of the observed fraction of hotspots/variants that occur on interaction interfaces over the fraction of interface residues on corresponding proteins (expected fraction). (E) Average number of protein interactions affected by hotspots and nonrecurrent variants. (F) Average edge betweenness of interactions affected by hotspots and nonrecurrent variants.











