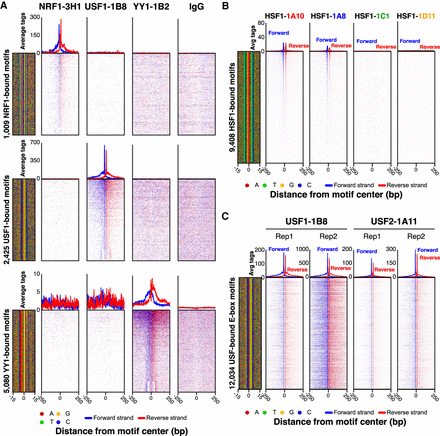
Validation of PCRP mAbs in ChIP-exo. (A) Comparison of ChIP-exo data at cognate versus noncognate motifs. ChIP-exo heatmap, composite, and DNA-sequence four-color plots were generated for NRF1, USF1, YY1, and IgG ChIP-exo data sets against the complete matrix of bound motifs from Supplemental Figure 2. The 5′ end of aligned sequence reads for each set of experiments was plotted relative to the distance from the cognate motif for each indicated target. Reads are strand-separated (blue, motif strand; red, opposite strand) and total-tag-normalized across samples. Rows are linked across samples and sorted based on their combined average rank-order in a 100-bp bin around each motif midpoint. High levels of background result in a more uniform distribution of reads across the window (as seen with the IgG control). (B) Enrichment comparisons of four unique HSF1 hybridoma clones (HSF1-1A10, HSF1-1A8, HSF1-1C1, HSF1-1D11). ChIP-exo heatmap, composite, and DNA-sequence four-color plots are shown for the indicated number and type of bound motifs for the indicated antibody hybridoma clones (A,B) or interaction partners (C) tested in K562 cells. The 5′ end of aligned sequence reads for each set of experiments was plotted against the distance from the cognate motif, present in the union of all called peaks between the data sets for each indicated target. Reads are strand-separated (blue, motif strand; red, opposite strand) and total-tag-normalized across samples. Rows are linked across samples and sorted based on their combined average rank-order in a 100-bp bin around each motif midpoint.











