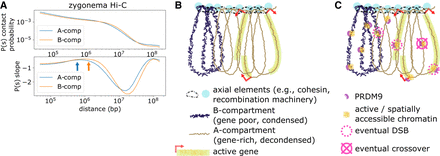
Proposed framework relating mammalian meiotic chromosomal architecture and recombination. (A) Contact probability versus genomic distance analysis of zygonema Hi-C data set. Orange and blue arrows indicate estimate of loop length for A- and B-compartments, respectively, determined as the maxima of the derivatives of the P(s) curves as in Gassler et al. (2017). (B) Simplified cartoon of proposed chromatin conformation. In early leptonema, meiotic chromosomes adopt a brush-loop architecture, with cohesin and recombination machinery located at the axis. Loops in the A-compartment have, on average, fewer base pairs than the B-compartment. Accordingly, A- and B-compartment regions depicted here represent roughly equal genomic lengths despite greater number of A-compartment loops. Physical size of A- and B-compartment loops may remain comparable owing to the relaxed linear packing density in A-compartment. (C) Concurrently during leptonema, PRDM9 binds across both A- and B-compartment regions, causing local increases in chromatin activity and spatial accessibility. Schematic depicts hypothetical example of 10 total binding events across the A- and B-compartment regions. After PRDM9 binding, a subset of binding loci are recruited to DSB machinery at the axis and form DSBs. This subset is biased toward A-compartment. Later during pachynema, a single crossover point is selected from the DSBs formed earlier, avoiding DSBs formed in gene-body regions.











