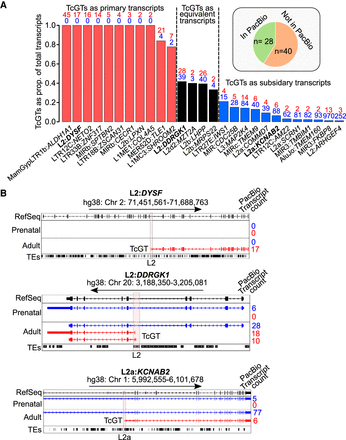
TcGTs are major contributors to neurodevelopmental transcript expression. (A, inset) Pie chart showing the number of TcGTs from Figure 4A detected with the same TE-derived TSS and isoform structure in PacBio long-read sequencing in the adult from Jeffries et al. (2020) in hg38. Bar charts show the proportion of total transcripts that are TcGT derived. Numbers above each bar represent the TcGT isoform PacBio transcript counts (red numbers) and annotated non-TcGT isoform PacBio transcript counts (blue numbers) in adult samples as determined by Jeffries et al. (2020). If some non-TcGT Ensembl-annotated isoforms had appreciably higher counts than others, only these were used. 5′-Truncated incomplete splice matches (as defined by Jeffries et al. 2020) for non-TcGTs were omitted unless similar in number to nontruncated transcripts. Red bars indicate the TcGT isoform is the primary transcript; black bars, TcGT isoforms have “equivalent” expression to canonical isoforms; and blue bars, TcGTs are subsidiary transcript isoforms. (B) Genome browser images of TcGTs (red) detected in long-read sequencing in prenatal and adult samples in hg38. Only the non-TcGT isoforms (blue) with the most PacBio transcript counts are shown for clarity. Vertical orange bars highlight the TE-derived TSS of the TcGTs and are the same as detected in our short-read RNA-seq analyses in hg19.











