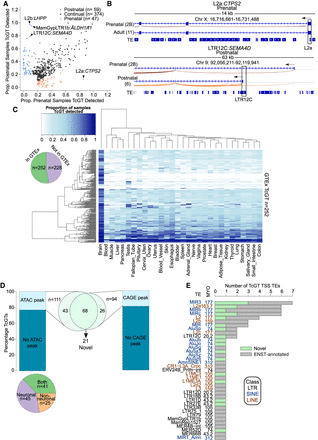
TE co-option as genic promoters drives spatiotemporal gene expression in human neurogenesis. (A) Dot plot showing the proportion of pre- or postnatal samples TcGTs were detected in and behaving similarly in both data sets (prenatal, postnatal, or continual). A TcGT was classed as “detected” if one or more reads were spliced between a TE and a genic exon. (B) Sashimi browser plots from the Integrative Genomics Viewer (IGV; Robinson et al. 2011; Thorvaldsdóttir et al. 2013) showing the splicing events in representative samples for prenatal enriched TcGT L2a:CTPS2 and the postnatal enriched LTR12C:SEMA4D. (C) Heatmap indicating the proportion of samples per GTEx tissue in which each TcGT from A was detected. Each row represents an individual TcGT and each column a different tissue. (C, inset) Pie chart indicating the proportion of neurodevelopmental TcGTs detected in GTEx. (D) Stacked barplots indicating the proportion of TcGT TE TSS loci overlapping an ATAC-seq peak from BOCA (left) and CAGE-peak from FANTOM5 (right), and pie charts indicating their cell type distribution (bottom left); ATAC and CAGE peak overlaps (center) and highlighting 21 novel, non-Ensembl-annotated transcripts. (E) Stacked barplots indicating the TE subfamily, TE class, TE age, and the Ensembl overlap of each TcGT TE TSS loci. For all TcGT information, see also Supplemental Table S8.











