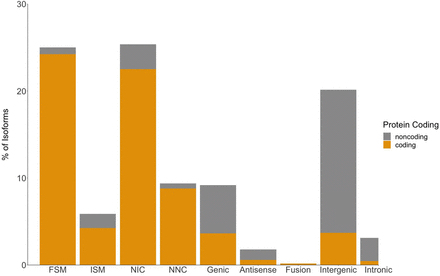
In-depth characterization of isoforms by SQANTI for Iso-Seq transcriptome. Isoforms are classified into nine different splice categories by SQANTI: full-splice match (FSM), incomplete splice match (ISM), novel in catalog (NIC), novel not in catalog (NNC), genic, antisense, fusion, intergenic, and intronic. Each splice category is divided into predicted protein-coding isoforms (gold) and predicted non-protein-coding isoforms (gray).











