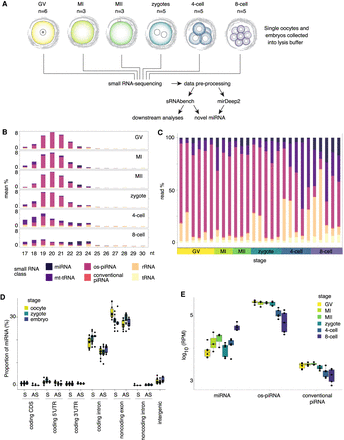
Categories of sRNAs. (A) Schematic of the study design. We sequenced sRNAs in human oocytes, zygotes, and embryos and did downstream analyses using sRNAbench (Aparicio-Puerta et al. 2019; Rueda et al. 2015) and miRDeep2 v0.1.2 (Friedländer et al. 2012). (GV) Germinal vesicle oocyte (n = 6); (MI) Metaphase I oocyte (n = 3); (MII) Metaphase II oocyte (n = 3); zygote (n = 5); (4-cell) four-cell embryo (n = 5); (8-cell) eight-cell embryo (n = 5). (B) sRNA class expression at different nucleotide lengths (17–30 nt) in the oocyte maturation stages, zygotes, and cleavage stage embryos. Stagewise mean proportion of reads mapped to different sRNA classes are shown. (miRNA) microRNA; (mt-tRNA) mitochondrial tRNA; (os-piRNA) oocyte short piRNA; (conventional piRNA) Piwi-interacting RNA; (rRNA) ribosomal RNA; (tRNA) transfer RNA. (C) Proportion of different sRNAs class reads in samples. The samples are grouped according to developmental stage and separated by sample number. (D) Proportion of miRNAs derived from different genomic elements at each developmental stage. (E) miRNA, os-piRNA, and conventional piRNA expression (reads per million [RPM]) in early human developmental stages. Embryos (n = 10) differed in os-piRNA expression from oocytes (n = 12) and zygotes (n = 5), and in conventional piRNA expression from oocytes (FDR < 0.05, Wilcoxon rank-sum test, two-sided), whereas for miRNAs the expression difference between oocytes and embryos was not significant (FDR = 0.054). (D,E) The horizontal line in the box plot indicates the median value.











