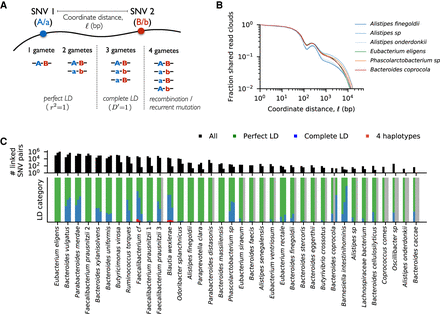
High levels of linkage disequilibrium (LD) in many resident populations. (A) Schematic of read cloud sharing between two SNVs separated by coordinate distance ℓ on the same reference contig. Three or fewer haplotypes are consistent with clonal evolution, whereas four haplotypes indicate a possible recombination event. (B) Observed fraction of shared read clouds as a function of ℓ for SNVs in the six example species in Figure 3. (C) Linkage disequilibrium between pairs of SNVs across a range of different species. The top panel shows the total number of linked SNV pairs (i.e., those with significantly elevated levels of read cloud sharing) for species in Figure 2 with sufficient coverage (Methods). For each species, the three bars denote SNV pairs with ℓ < 200 bp, 200 bp < ℓ < 2 kb, and ℓ > 2 kb, respectively. SNVs are included only if the minor allele has frequency f > 0.1. The bottom panel shows the observed proportion of SNV pairs in the top panel that fall in each of the LD categories illustrated in panel A. Across species, only a small fraction of SNV pairs provide evidence for recombination.











