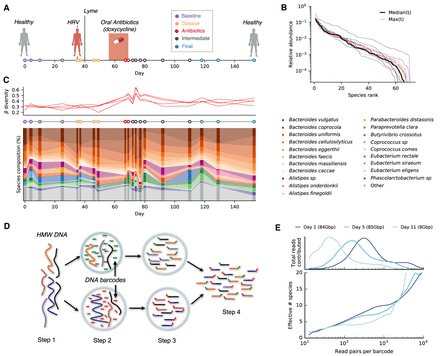
Read cloud sequencing of the gut microbiome of a single individual during disease, antibiotic treatment, and recovery. (A) Study design. Linked-read metagenomic sequencing was performed on 19 fecal samples collected from a single individual over a period of 5 mo. During this time, the individual was diagnosed with human rhinovirus (HRV) and Lyme disease and received an oral course of doxycycline. Samples are colored according to the epochs defined in Supplemental Table S1. (B) Rank relative abundance distribution at the species level, estimated from shotgun metagenomic reads (Methods). Colored lines show distributions obtained from individual time points, using the same color scheme as panel A. Solid and dashed black lines denote median and maximum relative abundances across all time points, respectively. (C) Species-level composition over time. Top panel illustrates Jensen-Shannon distance to each of four baseline samples as a function of time. Bottom panel shows relative abundance trajectories for a subset of the most abundant species; others are grouped together into the “other” category. (D) Schematic of linked-read sequencing with the 10x Genomics platform. High molecular weight metagenomic DNA is partitioned into millions of microfluidic droplets. Amplification and ligation reactions are performed within each droplet, yielding millions of short-read libraries that are tagged with droplet-specific DNA barcodes. The resulting “read clouds” are then pooled together and sequenced on an Illumina instrument. (E) Observed statistics of read clouds from the first three time points. The top panel shows total number of read pairs contributed by read clouds as a function of the number of read pairs they contain. The bottom panel shows a measure of the effective number of species that are detected in each read cloud as a function of the number of read pairs it contains (Methods). Many read clouds contain fragments from several different DNA molecules.











