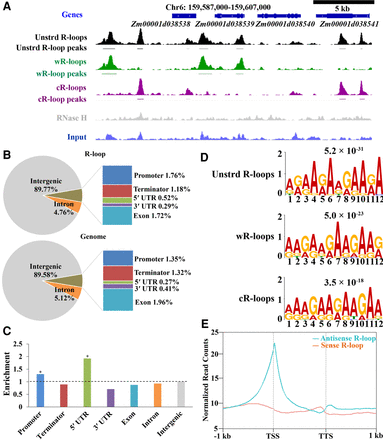
Genome-wide detection of R-loops in maize by ssDRIP-seq. (A) A representative region from Chromosome 6 showing the ssDRIP-seq data. Line 1, gene annotations; line 2, unstranded (unstrd) R-loop signal, normalized to the genome-wide mean; line 3, unstrd R-loop peaks; line 4, normalized wR-loops signal (green); line 5, wR-loop peaks; line 6, normalized cR-loop signal (purple); line 7, cR-loops peaks; line 8, DRIP signal of RNase H treatment (gray), normalized to the genome-wide mean of unstrd R-loops; line 9, input, normalized to the genome-wide mean. (B) Location analysis of unstranded R-loop peaks (upper) compared with the expected genomic distribution (lower) on the chromosome. The maize genome was characterized into seven classes that included six classes of genic regions (promoter, 5′ UTR, 3′ UTR, coding exon, intron, and terminator) and intergenic regions. Promoter, −1 kb to 100 bp of TSS; terminator, −100 bp to 1 kb of TTS. (C) Relative enrichment of unstranded R-loop peaks in different genomic elements. The dashed line represents enrichment fold = 1.0. Permutation test; asterisks denote the observed value was >90% of the permutation value. (D) DNA motif in the peak regions of unstranded R-loops, wR-loops, and cR-loops that were identified by MEME-ChIP. E-values are provided at the top. (E) Metaplots of sense (orange) and antisense (cyan) R-loop peaks centered on non-TE genes (B73_RefGen_v4), 95% mean.











