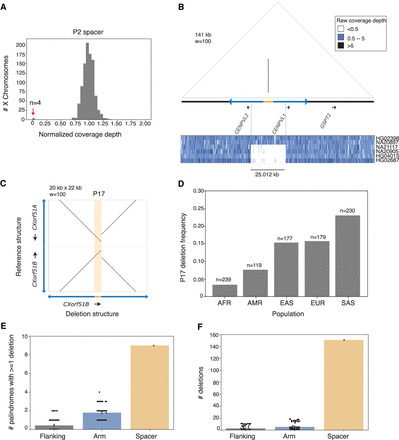
Deletions are enriched in human X-Chromosome spacers. (A) Normalized coverage depths for P2 spacer. Red arrow indicates four X Chromosomes with depth near 0. (B) Coverage depths across P2 and flanking sequence for two individuals with reference structure (HG02398, NA20897) and four with spacer deletions (NA21117, NA20905, HG04015, HG02687). (C) Square dot plot comparing palindrome centers (spacer + 10 kb inner arm on each side) for P17 reference structure and P17 deletion. (D) Frequency of P17 spacer deletions across five superpopulations from 1000 Genomes. (EUR) European, (AFR) African, (AMR) Admixed Americas, (EAS) East Asian, (SAS) South Asian. (E,F) Frequency of deletions detected in palindrome spacers compared to palindrome arms and flanking sequence. Size-matched regions from palindrome arms and single-copy sequence were selected at random; results from 100 iterations are shown.











