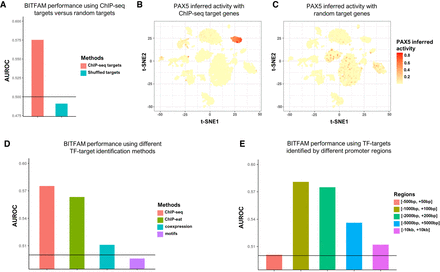
Figure 5.
Performance of BITFAM when prior knowledge is varied. (A) The AUROC of transcription factor activities by shuffling ChIP-seq data. (B) The inferred activities of PAX5 in Tabula Muris lung data by ChIP-seq target genes. (C) The inferred activities of PAX5 in Tabula Muris lung data by shuffled target genes. (D) AUROC of transcription factor targets identification methods on CRISPRi data set. (E) AUROC of promoter region on CRISPRi data set.











