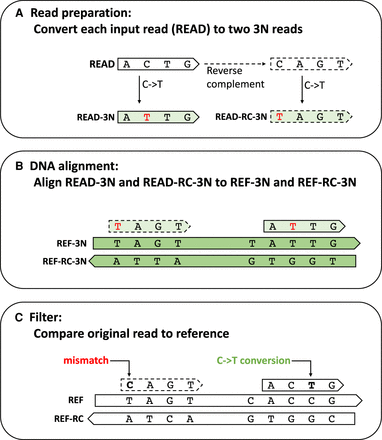
HISAT-3N alignment steps for BS-seq reads. (A) HISAT-3N converts each input read (READ) to two 3N reads: READ-3N and READ-RC-3N. READ-3N is READ with all thymine replaced by cytosine. READ-RC-3N is the reverse complement of READ, plus the replacement of cytosine with thymine. (B) HISAT-3N maps the two 3N reads to both REF-3N and REF-RC-3N references using prebuilt indexes. (C) After the 3-nt alignment, HISAT-3N compares the original read sequence (READ) to the original 4-nt references (REF and REF-RC) to identify unmethylated cytosine positions and recalculate an alignment score accordingly.











