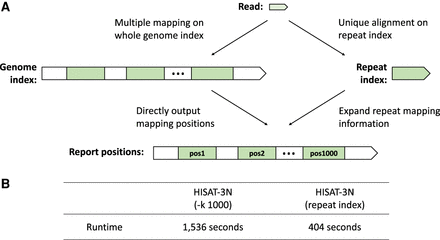
Repeat index enables faster 3-nt read alignment. (A) HISAT-3N aligns reads using two different strategies: (1) HISAT-3N can directly align reads to the whole genome using the genome index and output their mapped locations (A, left), and (2) HISAT-3N can use a repeat index to uniquely align reads to the repeat sequences regardless of how many locations to which they align on the genome (A, right). (B) Runtime comparison between direct mapping and repeat mapping strategy. The test data are 10 million simulated single-end BS-seq reads (0.2% per-base sequencing error rate).











