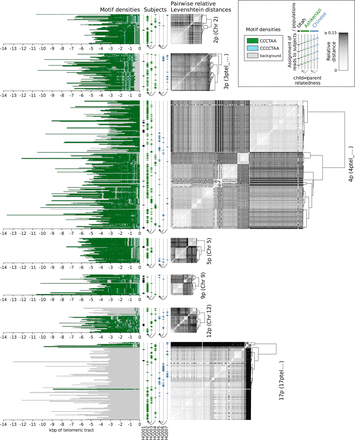
Clustering of reads by relative pairwise Levenshtein distances (unitless measure) on each chromosomal p arm of data sets HG001 through HG007, as well as densities of the top enriched motifs along each read. Each horizontal line represents an individual read; genomic coordinates are given in kilobase pairs, relative to the positions of the telomeric tract boundaries. Only the chromosomal arms cumulatively covered by at least 25 reads are displayed.











