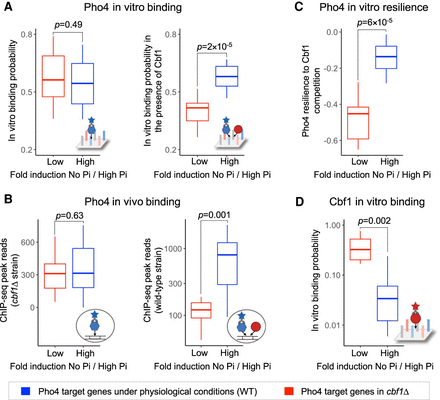
TF-TF competition contributes to differential gene activation. Box plots show the in vitro and in vivo TF binding data for sets of genes with low versus high fold induction in response to phosphate starvation (No Pi). Blue: Genes that are activated by Pho4 under physiological conditions, that is, in a wild-type strain where Cbf1 is present at physiological levels (Zhou and O'Shea 2011). Red: Genes that are activated by Pho4 only when Cbf1 is absent from the cell, that is, in a cbf1Δ strain (Zhou and O'Shea 2011). The two sets of Pho4 target genes were compared in terms of: (A) Pho4 in vitro binding (at 2 μM concentration) in the absence of Cbf1 (left) and in the presence of Cbf1 at 2 μM concentration (right), as measured by PBM and competition PBM, respectively; (B) Pho4 in vivo binding, as measured by ChIP-seq, in the absence of Cbf1 (left, cbf1Δ strain) and in the presence of Cbf1 (right, wild-type strain); (C) Pho4's in vitro resilience to Cbf1 competition (computed between competition PBM conditions: 2 μM Pho4 + 2 μM Cbf1 versus 2 μM Pho4 + 0.05 μM Cbf1) (Methods); and (D) Cbf1 in vitro binding (at 2 μM concentration), as measured by PBM. In vitro binding probabilities were computed from PBM or competition PBM data (Methods). In vivo binding levels are shown as read counts computed for Pho4 ChIP-seq peaks (Zhou and O'Shea 2011).











