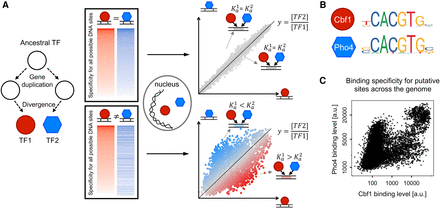
Paralogous TFs compete for DNA binding. (A) Schematic showing different TF-TF competition scenarios. If paralogous TFs have identical DNA-binding preferences, then their binding is determined by the levels of the TFs in the nucleus (top panels). However, most paralogs have diverged in specificity, binding differently at different sites even in the absence of other proteins (Berger et al. 2008; Badis et al. 2009; Wei et al. 2010; Gordân et al. 2013; Shen et al. 2018); this divergence leads to complex patterns of competitive binding, which depend on the TFs’ affinities and concentrations (bottom panels). (B) S. cerevisiae proteins Cbf1 and Pho4 s have similar, although not identical, DNA-binding specificities, as reflected by their position weight matrix (PWM) models (Sandelin et al. 2004). (C) Direct comparison between the in vitro binding levels of Cbf1 and Pho4 at their putative genomic binding sites, measured by genomic-context PBM (Gordân et al. 2013). Each data point corresponds to a 36-bp genomic region centered on a CACGTG site. Plot shows the fluorescence intensities from PBM assays, which are proportional to the level of bound TF at each genomic site. We note that fluorescence intensities are generally not directly comparable between PBM experiments for different proteins (see Supplemental Discussion). However, for all proteins tested, here and in prior studies (Berger et al. 2006; Siggers et al. 2011; Shen et al. 2018; Afek et al. 2020), the PBM fluorescence intensities correlate quantitatively with binding energies and equilibrium dissociation constants.











