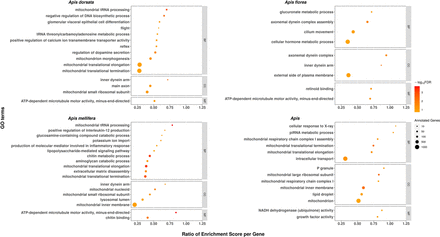
Functional categories enriched with genes under positive selection in each honey bee species and their most recent common ancestor. GO terms enriched in positively selected genes are depicted as spheres representing the number of annotated genes (sphere size) and the −log10 of their FDR (color intensity). GO enrichment scores, normalized by the number of annotated genes, are indicated by the x-axis. Most enriched GO terms with positively selected genes can be interpreted as adaptations to long-distance migration and increased colony size in A. dorsata, colony defense in A. florea, immunity in A. mellifera, and TE silencing and high recombination rates in the basal Apis lineage. (BP) Biological process; (CC) cellular component; (MF) molecular function.











