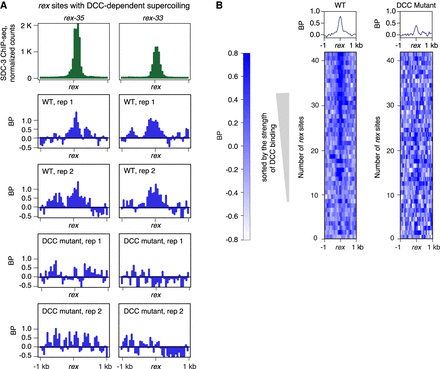
Local DCC-dependent negative supercoils occur at DCC binding sites (rex sites) with the highest DCC occupancy. (A) Examples of rex sites (rex-35 and rex-33) that show DCC-dependent negative supercoils. DCC subunit SDC-3 (green ChIP-seq signal) binds with high affinity to these rex sites. BP-seq signal (blue) is plotted for two replicates at each rex site in wild-type and DCC mutant embryos. Reduction of BP-seq signal in mutants shows that supercoiling is DCC dependent. These rex sites are at TAD boundaries on X. (B) Heatmaps of supercoils (BP-seq) are plotted in 2-kb regions around rex sites at 50-bp resolution in wild-type (replicate 1) and DCC mutant (replicate 1) embryos. rex sites were sorted according to their SDC-3 occupancy in wild-type embryos. Negative supercoiling correlates with strength of DCC occupancy and is eliminated in mutants (sdc-2) that prevent DCC binding.











