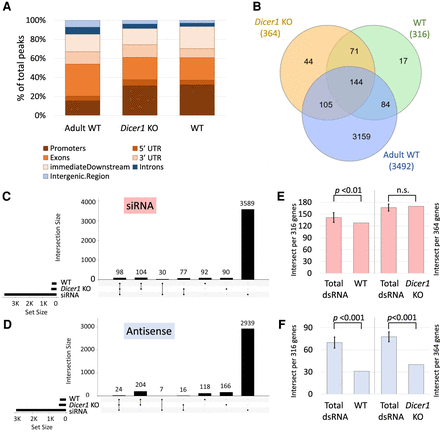
dsRNA formation in juvenile Dicer1 KO mice and age-matched wild-type controls. (A) dsRNA peaks in the three samples (Adult WT, Dicer1 KO, and WT) were annotated to biotypes using ChIPpeakAnno: Promoter, 5′ UTR, exon, 3′ UTR, immediate downstream (1000 bp) are in shades of brown, and introns and intergenic regions are in shades of blue. (B) Venn diagram depicting the genes associated with dsRNA peaks in Dicer1 KO, WT, and adult mouse testis and the overlaps between the samples. (C) The coordinates of genes associated with peaks in Dicer1 KO samples and WT controls were intersected with the coordinates of siRNA-associated genes (in red) (Hilz et al. 2017) and antisense genes (in blue) in D, respectively, and visualized in upset plots. In both cases, the Dicer1 KO specific genes show greater association with siRNA-associated and antisense genes (siRNA 77 vs. 30, antisense 16 vs. 7). (E) Number of dsRNA-associated genes in Dicer1 KO and WT controls that intersect with siRNA-associated genes, compared to an average of 10 size-matched random samples from the total dsRNA gene set. (F) Number of dsRNA-associated genes in Dicer1 KO and WT controls that intersect with antisense transcripts compared to the control total dsRNA gene set. Significance was determined by one-sample t-test.











