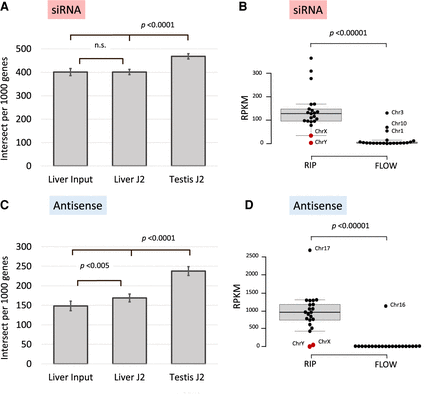
Reads in J2-enriched samples that intersect with endo-siRNAs (Hilz et al. 2017) and antisense transcripts. Samples from testis and liver as well as the flowthrough (FLOW) were analyzed, the upper panel (A,B) showing the overlap with siRNA forming genomic regions, the lower panel (C,D) with antisense genes. (A) Number of intersected genes per 1000 genes, dsRNA samples from testis show significantly more genes that generate endo-siRNAs than both J2-enriched samples and input control from liver. (B) Normalized number of fragments on individual chromosomes per megabase in combined RIP samples from mouse testis as compared to FLOW samples. Each dot represents a specific chromosome; the box plot indicates the median, 25th, and 75th percentiles as box limits and 1.5 times the interquartile range (whiskers); Chromosomes 1, 3, and 10 are clear outliers in the FLOW samples (Supplemental Fig. S3). The sex chromosomes tend to show a lesser overlap between dsRNA and endo-siRNA formation even if the lower number of dsRNA peaks is considered. (C) Number of dsRNA-forming genes intersected with antisense genes. The coordinates of 2991 antisense genes were retrieved and lists of 1000 randomly selected genes were intersected with lists from testis (1000 of 3492) and liver (1000 of 4900 J2 and 1000 of 4061 input control). The dsRNA-associated genes in both testis and liver are significantly associated with antisense genes (P < 0.0001 and P < 0.005, respectively). (D) Total number of reads on individual chromosomes per megabase in combined RIP and FLOW samples that intersect with antisense transcripts. The box plot gives the median, 25th, and 75th percentiles as box limits and 1.5 times the interquartile range (whiskers); Chromosome 16 represents an outlier in the FLOW samples.











