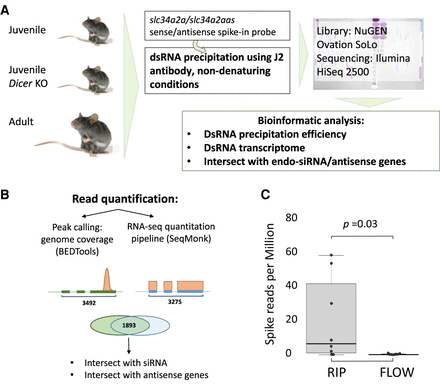
Experimental strategy to characterize the dsRNA transcriptome of mouse testis. (A) Testes from three groups of animals were collected, including 18-d-old wild-type and Dicer1 knockout mice as well as adult mice (4–6 mo old). The dsRNA was immune purified using the specific J2 antibody and both bound and unbound RNA samples were sequenced. (B) Two quantification methods were applied to call expressed genes; (1) peak calling using BEDTools (Quinlan and Hall 2010) followed by quantification using DESeq2, and (2) the RNA-seq pipeline in SeqMonk (https://www.bioinformatics.babraham.ac.uk/projects/seqmonk/) that quantifies exon mapping reads and returns RPKM values. Both pipelines yield lists of gene coordinates in BED/bedGraph format. (C) Enrichment of the spike-in probe in the J2 treated samples. Reads per million of the spike-in probe in the J2 immuno-enriched samples (RIP) versus the flowthrough samples (FLOW). The dots represent results from the eight different samples. t-test confirmed a significant accumulation of dsRNA in the J2-enriched samples.











