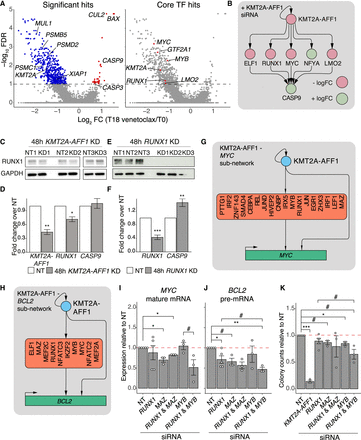
KMT2A-AFF1 and intermediate TFs cooperate to regulate cascade and FFL motif targets. (A) MAGeCK analysis of CRISPR screen comparing T0 (baseline) and T18 (venetoclax), plotting −log10 FDR against log2 gRNA FC. (Left) Key genes; (right) core GRN TFs. (B) Select cascade motifs in the KMT2A-AFF1 GRN that explain interactions between KMT2A-AFF1 and CASP9. Node color represents logFC response to KMT2A-AFF1 KD. (C,E) Western blot in SEM cells showing RUNX1 protein levels after 48 h KMT2A-AFF1 KD (C) or 48 h RUNX1 KD (E), with GAPDH as a loading control. (D,F) qRT-PCR assaying KMT2A-AFF1, RUNX1 and CASP9 expression following 48 h KMT2A-AFF1 KD (D) or 48 h RUNX1 KD (F) in SEM cells (n = 3). Expression normalized to GAPDH and shown relative to NT control. (G,H) Subnetworks illustrating interactions from KMT2A-AFF1 and cooperative TFs that feed into MYC (G) and BCL2 (H). (I,J) qRT-PCR analysis probing mature MYC mRNA (I) and BCL2 pre-mRNA (J) after 96 h KD targeting genes as indicated (n = 3, n = 5 for NT and RUNX1 KD). Expression normalized to GAPDH mature mRNA levels and shown relative to NT control. (K) Colony assay counts after 96 h KD targeting genes as indicated (n = 3, n = 5 for NT and RUNX1 KD). Colony counts shown relative to NT control. Error bars represent standard error of the mean; (#) P < 0.1; (*) P < 0.05; (**) P < 0.01; (***) P < 0.001.











