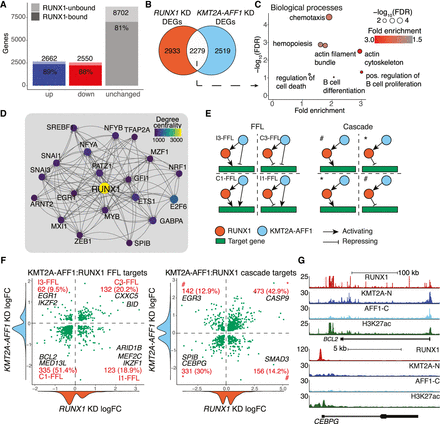
KMT2A-AFF1 cooperates with RUNX1 in FFL and cascade motifs to regulate downstream targets. (A) DEGs from nascent RNA-seq after 96 h RUNX1 KD. DEGs are defined as FDR < 0.05 (n = 3). Shaded area represents RUNX1-bound genes. (B) Overlap between KMT2A-AFF1 KD DEGs (Fig. 1C) and RUNX1 KD DEGs (A). (C) GO biological process enrichment for overlap shown in B. Size of points is proportional to significance. (D) The top 20 genes of the RUNX1 GRN by degree centrality. Lines indicate predicted interaction from protein to gene locus, with arrowheads pointing downstream. (E) FFL (left) and cascade (right) motifs. FFLs are subcategorized into C1-FFL, C3-FFL, I1-FFL, and I3-FFL as indicated. Cascade motifs are grouped into same sign of effect (*) and opposing sign of effect (#). (F) Scatter plot of RUNX1 and KMT2A-AFF1 KD logFC response at FFL (left) and TF cascade (right) target genes. Density plots along the axis show distribution of logFC values. Quadrants of scatter plots align with FFL and cascade types shown in E. (G) ChIP-seq tracks for KMT2A-N, AFF1-C, RUNX1, and H3K27ac, normalized to 1 × 107 reads. (Top) BCL2, a FFL target, bound by both KMT2A-AFF1 and RUNX1. (Bottom) CEBPG, a cascade target, bound by RUNX1 only.











