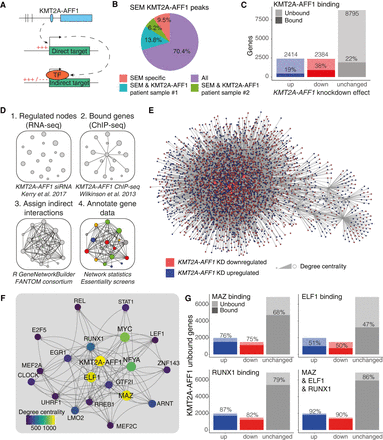
Developing a GRN model to assess regulatory impact of KMT2A-AFF1. (A) Schematic illustrating the concept of KMT2A-AFF1-targeted TFs, leading to indirect downstream regulation. (B) Proportion of KMT2A-AFF1-bound genes in SEM cells (nearest annotated promoter in ChIP-seq) that overlap with KMT2A-AFF1 targets from patient samples. (C) DEGs from nascent RNA-seq after 96 h KMT2A-AFF1 KD. DEGs are defined as FDR < 0.05 (n = 3). Shaded area represents KMT2A-AFF1-bound genes. (D) GRN creation workflow using nascent RNA-seq and ChIP-seq data (Methods). (E) Visualization of the whole network. Node color represents down-regulation (red) and up-regulation (blue) upon KMT2A-AFF1 KD. (F) The top 20 genes of the KMT2A-AFF1 GRN by degree centrality. Lines indicate predicted interaction from protein to gene locus, with arrowheads pointing downstream. (G) KMT2A-AFF1 KD DEGs that are unbound by KMT2A-AFF1, as highlighted in C. Shaded areas represent MAZ, ELF1, or RUNX1-bound genes.











